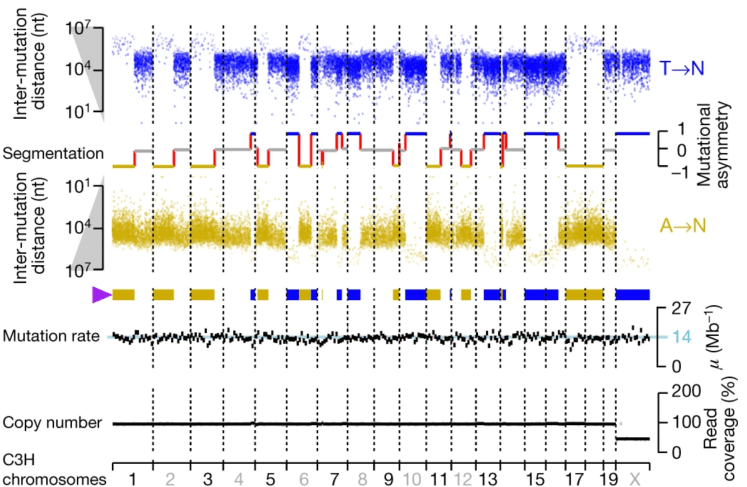
For almost 3 years we’ve had the pleasure of participating in the Liver Cancer Evolution (LCE) consortium, along with the labs of Duncan Odom, Paul Flicek, Nuria Lopez-Bigas and Martin Taylor – and today sees the publication of the first consortium paper in Nature. The Odom lab generated some unprecedented data, mapping genome and transcriptome evolution during liver tumour evolution in several strains/species of mice some time ago, and we have all collaborated on the analysis of these data. The initial analyses revealed surprising patterns of mutational asymmetry in these model systems: huge multimegabase segments of each chromosome showed strong biases to particular base substitutions. These patterns emerge due to a failure to repair mutagenic DNA lesions over successive cell cycles, and similar patterns can appear in human cells following mutagenesis. Each new round of DNA replication on a lesion-containing strand can lead to the incorporation of a different mispaired base opposite the lesion site in the newly synthesized strand, generating cells in the evolving tumour with different mutations of the same base pair. Unrepaired lesion segregation is therefore an unexpected source of diversity during what is otherwise straightforward clonal evolution, and may provide fuel for adaptive evolution in early tumours.